Medical Image Segmentation Training Pipeline
About This Architecture
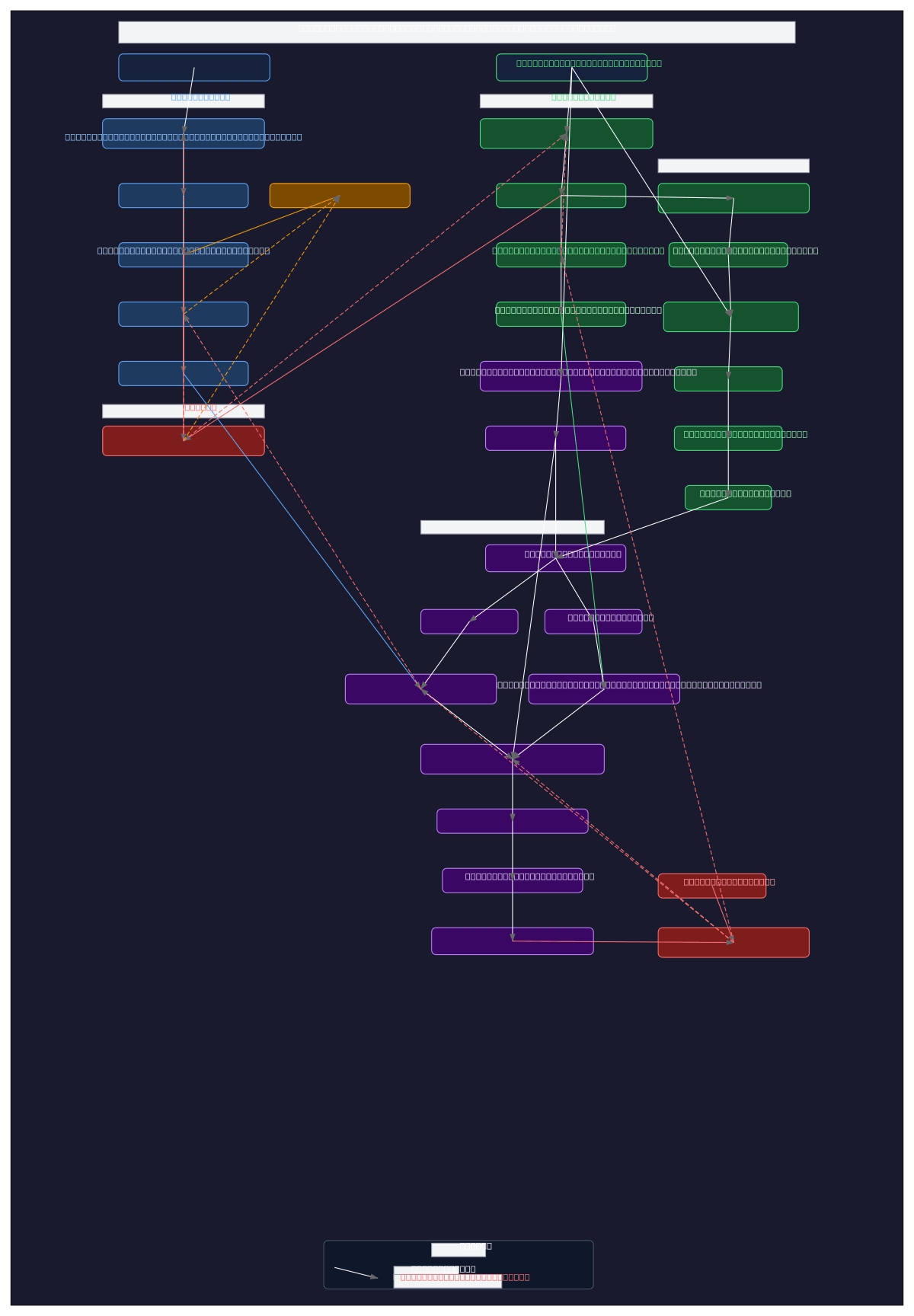
Prompt-based medical image segmentation training pipeline integrating BiomedCLIP encoders, SAM with LoRA adapters, and learnable prompt tokens for colonoscopy polyp detection. The architecture branches into text, image, and saliency pathways, with BiomedCLIP processing natural language prompts and colonoscopy images through partially unfrozen encoders, while SAM components receive enriched dense and sparse prompts via projection layers and context vectors. M2IB information bottleneck with learnable gates generates saliency heatmaps refined by differentiable CRF, feeding bounding box extraction into SAM's mask decoder for final segmentation. Training uses DHN-NCE contrastive loss on text embeddings and Dice loss on predicted masks against ground truth, enabling efficient fine-tuning of foundation models for clinical image analysis. Fork and customize this pipeline on Diagrams.so to adapt prompt engineering strategies, adjust LoRA ranks, or integrate alternative vision encoders for your medical imaging tasks.
People also ask
How do you build a medical image segmentation model that accepts natural language prompts and integrates vision-language foundation models like BiomedCLIP with SAM?
This diagram shows a complete training pipeline where text prompts are encoded by BiomedCLIP's text encoder into embeddings enriched with learnable context vectors, while colonoscopy images flow through BiomedCLIP's vision encoder and M2IB information bottleneck to generate saliency heatmaps. Both pathways feed into SAM's prompt encoder and mask decoder (with LoRA adapters for efficient fine-tunin
- Domain:
- Ml Pipeline
- Audience:
- Machine learning engineers building medical image segmentation models with vision-language foundation models
Generated by Diagrams.so — AI architecture diagram generator with native Draw.io output. Fork this diagram, remix it, or download as .drawio, PNG, or SVG.