U-Net DFE-ASPP Segmentation Architecture
About This Architecture
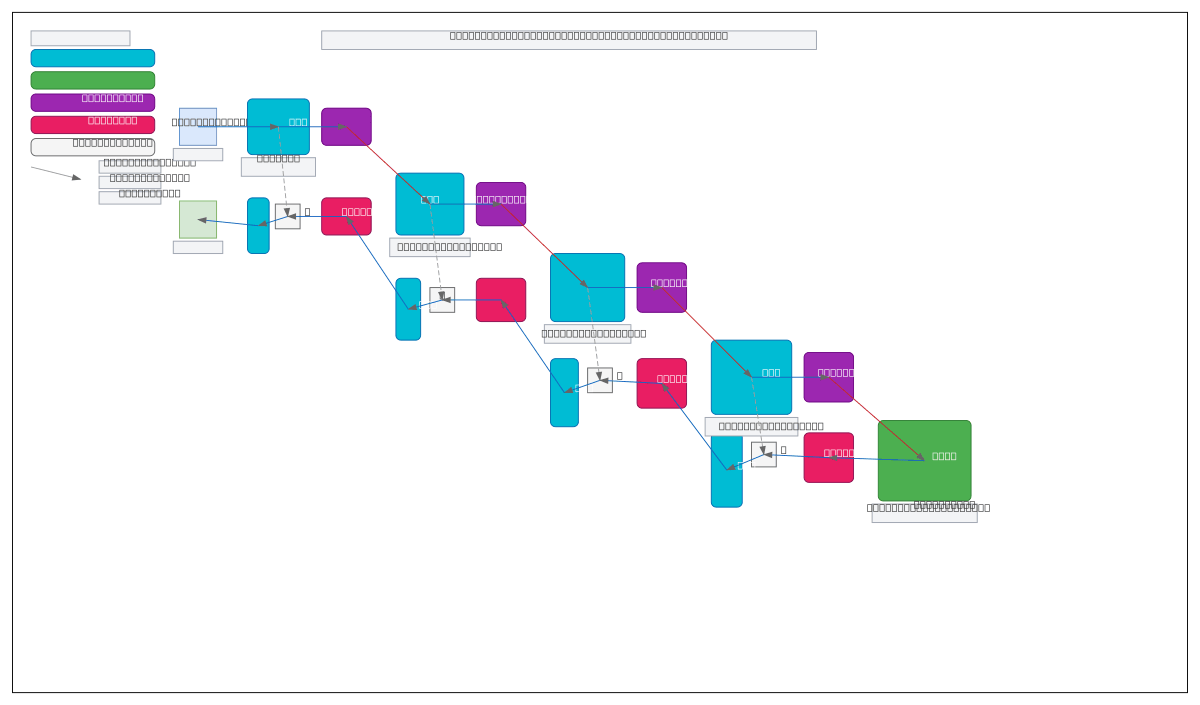
U-Net with Deformable Feature Enhancement (DFE) and Atrous Spatial Pyramid Pooling (ASPP) combines multi-scale feature extraction with adaptive convolutions for precise medical image segmentation. Brain MRI input flows through four downsampling stages with DFE modules, converges at an ASPP bottleneck, then reconstructs via four upsampling stages with concatenated skip connections and DFE refinement. This architecture captures both fine anatomical details and global context, critical for accurate tumor, lesion, and organ segmentation in clinical workflows. Fork and customize this diagram on Diagrams.so to document your own segmentation pipeline or compare encoder-decoder variants. The DFE modules enable the network to learn spatially adaptive receptive fields, improving robustness to anatomical variations across patient populations.
People also ask
How does U-Net with deformable convolutions and ASPP improve medical image segmentation accuracy?
This diagram shows how DFE modules enable spatially adaptive feature learning across four downsampling stages, while ASPP at the bottleneck captures multi-scale context via atrous convolutions. Skip connections preserve fine anatomical details during upsampling, and DFE refinement in the decoder adapts features to local image content, improving segmentation precision for brain tumors and lesions.
- Domain:
- Ml Pipeline
- Audience:
- Machine learning engineers and medical imaging researchers implementing deep learning segmentation models
Generated by Diagrams.so — AI architecture diagram generator with native Draw.io output. Fork this diagram, remix it, or download as .drawio, PNG, or SVG.