SAM Polyp Segmentation Pipeline
About This Architecture
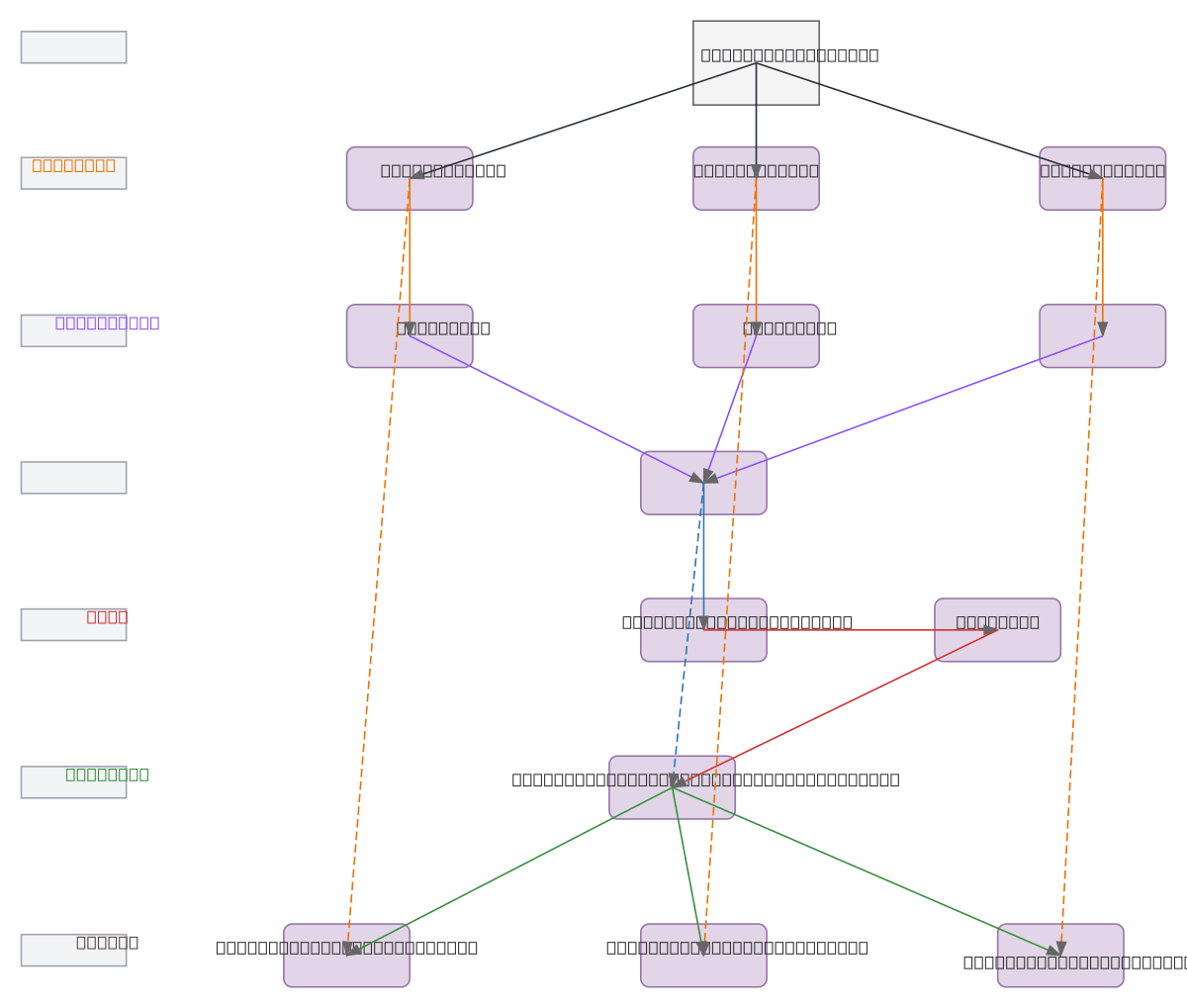
SAM Polyp Segmentation Pipeline uses three Vision Transformer encoders (ViT1, ViT2, ViT3) to process polyp images in parallel, each feeding through dedicated adapters into a unified Feature Fusion module. The fused features flow through an Edge Extraction Module to generate edge maps, which are combined with the original fused features via residual addition to create edge-enhanced outputs. This multi-encoder architecture with edge-aware residual connections improves segmentation accuracy by capturing both semantic and boundary information across multiple transformer scales. Fork and customize this pipeline on Diagrams.so to adapt it for your specific endoscopy datasets or integrate it into your medical imaging workflow.
People also ask
How does the SAM polyp segmentation pipeline combine multiple Vision Transformers for accurate medical image analysis?
The SAM pipeline processes polyp images through three parallel ViT encoders (ViT1, ViT2, ViT3), each adapted via dedicated adapters and fused into a unified feature representation. Edge maps extracted from fused features are combined with residual connections to enhance boundary detection, improving segmentation precision for endoscopy applications.
- Domain:
- Ml Pipeline
- Audience:
- Computer vision engineers and medical imaging researchers implementing polyp segmentation models
Generated by Diagrams.so — AI architecture diagram generator with native Draw.io output. Fork this diagram, remix it, or download as .drawio, PNG, or SVG.