MSDCA-UNet Research Workflow Diagram
About This Architecture
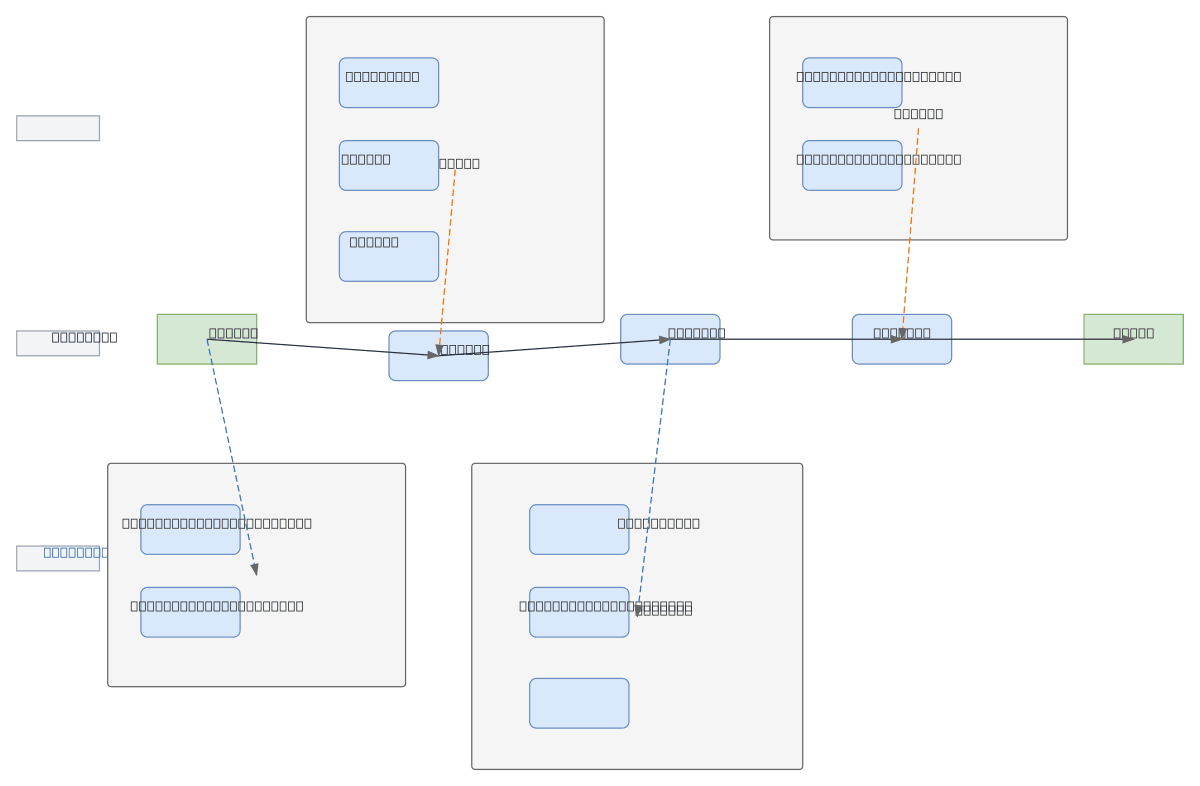
MSDCA-UNet is a multi-scale deep learning architecture for medical image segmentation that combines standard U-Net with deformable convolution and attention mechanisms to handle tumors with variable sizes and blurred infiltrative boundaries. The research workflow progresses from problem definition through model design (baseline U-Net enhanced with DSAM and MSCFM modules), training with joint loss functions and ablation studies, to system implementation with image loading, slice browsing, and mask rendering capabilities. Evaluation uses Dice coefficient, false positive analysis, and extreme slice visualization to validate performance on complex lesions. This architecture addresses key challenges in medical imaging where tumor scale variance and edge ambiguity demand multi-scale feature fusion and adaptive spatial attention. Fork and customize this workflow diagram on Diagrams.so to document your own segmentation research pipeline or integrate it into OCI-based medical AI platforms.
People also ask
How does MSDCA-UNet improve tumor segmentation for lesions with variable sizes and blurred boundaries?
MSDCA-UNet enhances standard U-Net by integrating deformable convolution (DSAM) and multi-scale feature fusion (MSCFM) modules to adaptively capture tumors across different scales and handle infiltrative edge ambiguity. The architecture uses joint loss functions and attention mechanisms validated through ablation studies and Dice coefficient evaluation.
- Domain:
- Ml Pipeline
- Audience:
- Machine learning researchers and medical imaging engineers developing deep learning segmentation models
Generated by Diagrams.so — AI architecture diagram generator with native Draw.io output. Fork this diagram, remix it, or download as .drawio, PNG, or SVG.